Step 9: Sequencing
Chromium Single Cell Multiome ATAC libraries comprise double stranded DNA with standard Illumina® paired-end constructs which begin with P5 and end with P7.
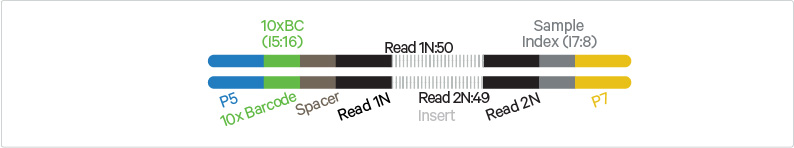
Sequencing these libraries produces a standard Illumina® BCL data output folder that includes paired-end Read 1N and Read 2N used for sequencing the DNA insert, along with the 8 bp sample index in the i7 read and 16 bp 10x Barcode sequence in the i5 read.
Chromium Single Cell Multiome ATAC Library
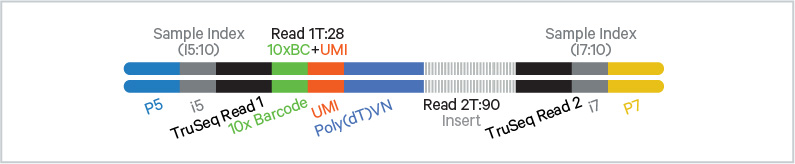
Chromium Single Cell Multiome Gene Expression libraries comprise cDNA insert with standard Illumina® paired-end constructs which begin with P5 and end with P7. Sequencing these libraries produces a standard Illumina® BCL data output folder.
TruSeq Read 1 is used to sequence 16 bp 10x Barcodes and 12 bp UMI, while 10 bp i5 and i7 sample index sequences are the sample index reads. TruSeq Read 2 is used to sequence the insert.
Chromium Single Cell Multiome Gene Expression Library