Step Overview (Step 8.1d)
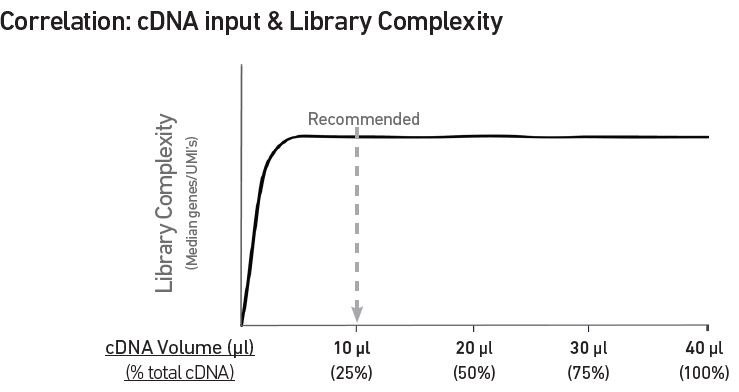
Correlation between input & library complexity
A Single Cell Gene Expression library is generated using a fixed proportion (10 μl, 25%) of the total cDNA (40 μl) obtained at step 7.2n. The complexity of this library will be comparable to one generated using a higher proportion (>25%) of the cDNA. The remaining proportion (30 μl, 75%) of the cDNA may be stored at 4°C for up to 72 h or at −20°C for longer-term storage (up to 4 weeks).
Note that irrespective of the total cDNA yield (ng), which may vary based on cell type, targeted nuclei recovery etc., this protocol has been optimized for a broad range of input mass (ng), as shown in the example below. The total number of SI PCR cycles (step 8.5d) should be optimized based on carrying forward a fixed proportion (10 μl, 25%) of the total cDNA yield calculated during Post cDNA Amplification QC & Quantification (step 7.3).
|
Cell Type |
Targeted Nuclei Recovery |
Total cDNA Yield (ng) |
cDNA Input into Fragmentation |
SI PCR |
|
|---|---|---|---|---|---|
|
Volume (µl) |
Mass (ng) |
||||
|
High RNA Content
|
|
150 ng |
10 µl |
37.5 ng |
14 |
|
|
400 ng |
10 µl |
100 ng |
13 |
|
|
Low RNA Content
|
|
1 ng |
10 µl |
0.25 ng |
16 |
|
|
100 ng |
10 µl |
25 ng |
14 |
|
The term "cell" as used here applies to nuclei too.





