7.3 cDNA QC & Quantification
-
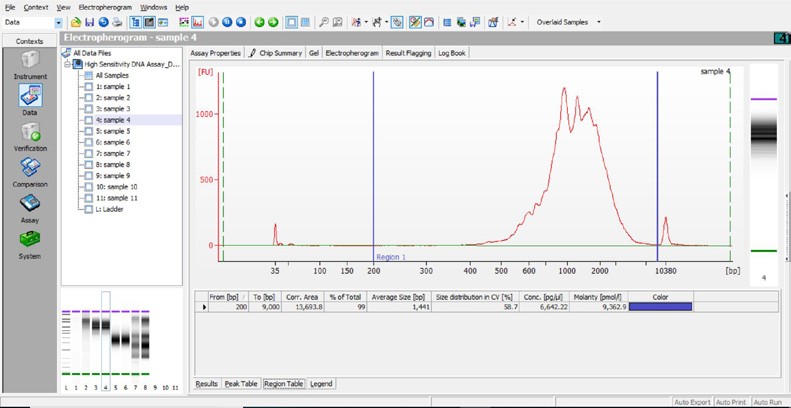
Run 1 μl undiluted sample on an Agilent Bioanalyzer High Sensitivity chip.
Depending on specific sample, it may also be diluted 1:2. Lower molecular weight product (35 – 150 bp) may be present on the traces. This is normal and does not affect sequencing or application performance.
|
EXAMPLE CALCULATION |
|
|---|---|
|
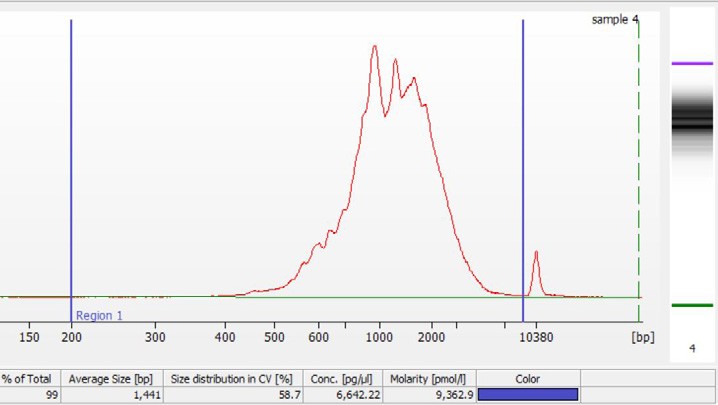
i. Select Region Under the “Electropherogram” view choose the “Region Table”. Manually select the region of ~200 – ~9000 bp
|
iii. Calculate Multiply the cDNA concentration [pg/µl] reported via the Agilent 2100 Expert Software by the elution volume (40 µl) of the Post cDNA Amplification Reaction Clean Up sample (taking any dilution factors into account) and then divide by 1000 to obtain the total cDNA yield in ng.
Example Calculation of cDNA Total Yield
Concentration: 6642.22 pg/µl Elution Volume: 40 Dilution Factor: 1 Total cDNA Yield = Conc’n (pg/µl) x Elution Volume (µl) x Dilution Factor 1000 (pg/ng)
= 6642.22 (pg/µl) x 40 (µl) x 1 = 265.69 ng 1000 (pg/ng)
Construction (step 8) = 0.25 x Total cDNA yield
= 0.25 x 265.69= 66.42 ng
Refer to step 8.5 for appropriate number of Sample Index PCR cycles based on carry forward cDNA yield/input cDNA. |
|
ii. Note Concentration [pg/µl]
|
|
Alternate QC Methods
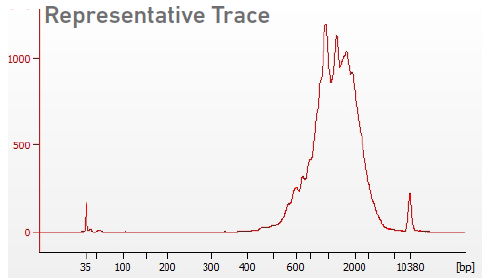
See Appendix for representative traces:
Agilent Bioanalyzer, Agilent TapeStation, or LabChip are the recommended methods for accurate quantification.